Code
install.packages("BiocManager")
BiocManager::install("SpatialArtifacts")SpatialArtifacts is an R package that provides a data-driven a two-step workflow to identify, classify, and handle spatial artifacts in spatial transcriptomics data from multiple platforms including 10x Visium (Standard and HD). Broadly, the idea behind the package is we that combine median absolute deviation (MAD)-based outlier detection with morphological image processing to identify the artifacts. These artifacts, often appearing as areas of low gene/UMI counts or high mitochondrial ratio at tissue edges (edge artifacts) or in the interior (interior artifacts), can negatively impact downstream analyses. The methods are implemented as an R package within the Bioconductor framework, and is available from Bioconductor.
In the following, we provide an overview of the functionality in the package and we demonstrate how to apply the package on real-world datasets across different spatial transcriptomics platforms.
More details describing the method are available in our paper, available from bioRxiv.
The following code will install the latest release version of the SpatialArtifacts package from Bioconductor. Additional details are shown on Bioconductor.
install.packages("BiocManager")
BiocManager::install("SpatialArtifacts")The latest development version can also be installed from the devel version of Bioconductor or from GitHub.
In the examples below, we assume the input data are provided as a SpatialExperiment Bioconductor object. In this case, the outputs are stored in the rowData of the SpatialExperiment object.
SpatialArtifacts is designed to work across multiple spatial transcriptomics platforms:
:::.::: {.callout-important}
The morphological detection framework automatically adapts to different grid arrangements, but parameter scaling is critical for optimal performance across platforms.
:::
The core philosophy is a two-step process: detect, and then classify. This separates the sensitive task of identifying all potential problem spots from the more nuanced task of deciding what to do with them.
In the first step, we use the detectEdgeArtifacts() function. Here, the goal is to identify all spots that could potentially be part of an artifact.
How does it work:
sum_gene) using a Median Absolute Deviation (MAD) threshold.focal_transformations():
my_fill): Fill spots completely surrounded by outliersmy_outline): Fill spots outlined by outliers in a larger 16-pixel perimetermy_fill_star): Fill spots with outliers in all four cardinal directions (N, S, E, W)min_cluster_size threshold (default: 40 spots)problemAreas)clumpEdges(). For each of the four borders (north, south, east, west), calculates the proportion of boundary spots belonging to each cluster. A cluster is classified as an edge artifact if this proportion meets or exceeds edge_threshold (default: 0.75, meaning ≥75% border coverage) on any single border directionThe output from this phase will add three raw columns to your spe object: _edge, _problem_id, and _problem_size
In the second step, we use the classifyEdgeArtifacts() function. Here the goal is to take the raw detections from the previous step and apply a clear, hierarchical logic to assign final labels.
How does it work:
spe object processed by detectEdgeArtifacts()_edge_artifact (TRUE or FALSE) based on the detection step?_problem_size larger than min_spots (default: 20)?"not_artifact" — High-quality spots"large_edge_artifact" — Large artifact cluster (> min_spots) touching the tissue edge"small_edge_artifact" — Small artifact cluster (≤ min_spots) touching the tissue edge"large_interior_artifact" — Large artifact cluster (> min_spots) located inside the tissue"small_interior_artifact" — Small artifact cluster (≤ min_spots) located inside the tissueThe output from this phase will add one classification column named _classification.
Tuning the parameters lets you adapt the workflow to different tissue types, data quality, and spatial transcriptomics platforms. The package uses a wrapper function that routes to platform-specific implementations.
CRITICAL FIRST STEP: Specify your platform using the platform parameter in detectEdgeArtifacts() function:
| Platform | Function Call | Required Parameters |
|---|---|---|
| Standard Visium | detectEdgeArtifacts(spe, platform="visium", ...) |
shifted (usually TRUE) |
| VisiumHD | detectEdgeArtifacts(spe, platform="visiumhd", resolution="16um", ...) |
resolution (“8um” or “16um”) |
# Standard Visium (55µm hexagonal grid)
spe <- detectEdgeArtifacts(spe, platform = "visium", shifted = TRUE, ...)
# VisiumHD 16µm (square grid)
spe <- detectEdgeArtifacts(spe, platform = "visiumhd", resolution = "16um", ...)
# VisiumHD 8µm (square grid)
spe <- detectEdgeArtifacts(spe, platform = "visiumhd", resolution = "8um", ...)detectEdgeArtifacts()
The wrapper function accepts platform-specific parameters that are routed to the appropriate implementation.
platform (REQUIRED) – Character string: "visium" or "visiumhd" (case insensitive)
qc_metric (Default: "sum_gene") – Column name for QC metric used in outlier detection
"sum_umi", "detected", "nFeature"
samples (Default: "sample_id") – Column name for sample identifiers
mad_threshold (Default: 3) – Sensitivity for detecting outliers
name (Default: "edge_artifact") – Prefix for output column names
[name]_edge, [name]_problem_id, [name]_problem_size
verbose (Default: TRUE) – Print progress messages
keep_intermediate (Default: FALSE) – Retain intermediate outlier detection columns
When platform = "visium", use:
edge_threshold (Default: 0.75) – Minimum proportion of a tissue boundary that must be occupied by outlier clusters (collectively) for those clusters to be classified as edge artifacts.
Important Behavior: The threshold is applied to the total coverage of all outlier clusters on each boundary direction (North, South, East, West). If multiple clusters collectively cover ≥ edge_threshold of a boundary, all clusters touching that boundary are classified as edge artifacts, even if no single cluster meets the threshold individually.
Example Scenario:
Rationale: Edge drying artifacts typically affect large, continuous regions along tissue boundaries. Multiple clusters on the same boundary often result from a single underlying technical failure (incomplete permeabilization) and should be treated as a unified artifact rather than independent events.
Tuning Guidance:
min_cluster_size (Default: 40) – Minimum cluster size (in spots) for morphological cleaning during focal transformation steps
shifted (Default: FALSE) – Apply coordinate adjustment for hexagonal grid alignment
shifted = TRUE when using array coordinates from Space Ranger.FALSE if using pre-transformed pixel coordinates or for VisiumHD square grids.batch_var (Default: "both") – Determines grouping for MAD calculation
"sample_id", "slide", or "both"
"both": Spots flagged as outliers if below threshold in either sample or slide groupingWhen platform = "visiumhd", use:
resolution (REQUIRED) – Character string: "8um" or "16um"
buffer_width_um (Default: 80) – Buffer zone width in micrometers (physical units)
resolution:
min_cluster_area_um2 (Default: 1280) – Minimum cluster area in square micrometers (physical units)
resolution:
col_x and col_y (Default: "array_col", "array_row") – Column names for bin coordinates
Key Difference from Visium: VisiumHD parameters are specified in physical units (µm, µm²) rather than bin counts. This ensures consistency across resolutions while the algorithm handles the bin conversion internally.
classifyEdgeArtifacts()
The classification step is platform-independent but requires appropriate parameter scaling.
min_spots (Default: 20) – CRITICAL PARAMETER: Threshold (in number of spots/bins) to distinguish "large" from "small" artifacts
Platform-Specific Scaling Required:
This parameter must be adjusted based on spatial resolution to represent equivalent physical artifact sizes:
| Platform | Recommended min_spots
|
Physical Area | Scaling Factor |
|---|---|---|---|
| Standard Visium (55µm) | 20-40 |
~0.06-0.12 mm² | Baseline (1×) |
| VisiumHD 16µm bins | 100-200 |
~0.026-0.051 mm² | ~6-10× Visium |
| VisiumHD 8µm bins | 400-800 |
~0.026-0.051 mm² | ~20-40× Visium |
Automatic Scaling Formula:
min_spots_HD <- min_spots_visium × (55 / bin_size_µm)²
# Example: For min_spots = 30 on Standard Visium
# VisiumHD 16µm: 30 × (55/16)² ≈ 354 bins
# VisiumHD 8µm: 30 × (55/8)² ≈ 1,420 binsWhy scaling matters: The same physical artifact (e.g., 0.1 mm² edge dryspot) will cover:
Without scaling, large VisiumHD artifacts would be incorrectly classified as “small.”
qc_metric (Default: "sum_umi") – QC metric column for validation (must exist but not directly used in classification logic)
samples (Default: "sample_id") – Sample ID column name
exclude_slides (Default: NULL) – Vector of slide IDs to exclude from edge classification
FALSE
name (Default: "edge_artifact") – Must match the name used in detectEdgeArtifacts()
| Feature | Standard Visium | VisiumHD |
|---|---|---|
| Grid Type | Hexagonal | Square |
Requires shifted? |
Yes (TRUE) |
No (not used) |
| Resolution Parameter | Not applicable |
Required ("8um" or "16um") |
| Edge Detection Method | Morphological + boundary coverage | Buffer zone + morphological |
| Parameter Units | Spot counts | Physical units (µm, µm²) |
Default min_spots (classify) |
20-40 | 100-200 (16µm), 400-800 (8µm) |
| Typical Dataset Size | ~5,000 spots | ~480k bins (16µm), ~1.9M bins (8µm) |
After both functions, several columns are added to colData(spe):
*_edge – Raw detection: Is the spot in a cluster touching the tissue border? (TRUE/FALSE)*_problem_id – Raw detection: ID of the problem area.*_problem_size – Raw detection: Size (number of spots) of the problem area.*_true_edges — Intermediate: Edge status after applying exclude_slides (used by classifyEdgeArtifacts()).*_classification — Final classification: One of "not_artifact", "large_edge_artifact", "small_edge_artifact", "large_interior_artifact", or "small_interior_artifact".This package includes spe_vignette, a lightweight SpatialExperiment object derived from a human hippocampus Visium sample.
This vignette will load this raw-like object and run the full SpatialArtifacts workflow on it live.
Note: To meet package size requirements (< 5MB), this object has been subset (e.g., to coding genes) and sparsified, but no artifact detection has been run. We will perform those steps now.
The underlying spatial clustering functions in this package currently require a dense matrix to perform coordinate-based calculations. We must first convert the sparse counts assay in our spe_vignette object to a standard (dense) matrix.
data(spe_vignette)
cat("Loaded data dimensions:", dim(spe_vignette), "\n")
#> Loaded data dimensions: 12971 4965
assay(spe_vignette, "counts") <- as.matrix(assay(spe_vignette, "counts"))
names(colData(spe_vignette))[names(colData(spe_vignette)) == "sum"] <- "sum_umi"
spe_detected <- detectEdgeArtifacts(
spe_vignette,
platform = "visium", # IMPORTANT: Specify Standard Visium platform
qc_metric = "sum_umi",
samples = "sample_id",
shifted = TRUE, # Hexagonal grid coordinate adjustment
batch_var = "sample_id",
mad_threshold = 3,
edge_threshold = 0.75,
name = "edge_artifact"
)
#> Detecting edges...
#> Sample V11L05-335_C1: 74 edge spots detected
#> Finding problem areas...
#> Removed intermediate columns: lg10_sum_umi, sum_umi_3MAD_outlier_sample, sum_umi_3MAD_outlier_binary
#> Edge artifact detection completed!
#> Total edge spots: 74
#> Total problem area spots: 78
cat("\n=== RESULTS ===\n")
#>
#> === RESULTS ===
table(Edge_Detected = spe_detected$edge_artifact_edge)
#> Edge_Detected
#> FALSE TRUE
#> 4891 74
# Classification with Standard Visium parameters
spe_classified <- classifyEdgeArtifacts(
spe_detected,
min_spots = 20,
name = "edge_artifact"
)
#> Classifying artifacts spots...
#> Classification added: edge_artifact_classification
#>
#> Classification summary:
#> not_artifact: 4887 spots
#> small_edge_artifact: 74 spots
#> small_interior_artifact: 4 spots
cat("\n=== Classification Results ===\n")
#>
#> === Classification Results ===
table(spe_classified$edge_artifact_classification)
#>
#> not_artifact small_edge_artifact small_interior_artifact
#> 4887 74 4For VisiumHD data, the workflow is identical except for parameter scaling. Here’s a complete example showing how to adapt parameters for VisiumHD:
# This is a pseudo-example demonstrating VisiumHD 16µm workflow
# Assumes you have loaded a VisiumHD SpatialExperiment object as 'spe_hd16'
# Step 1: Ensure required QC metrics are calculated
library(scuttle)
spe_hd16 <- addPerCellQCMetrics(spe_hd16)
# Step 2: Detection Phase - VisiumHD uses square grid (no 'shifted' needed)
spe_hd16_detected <- detectEdgeArtifacts(
spe_hd16,
platform = "visiumhd", # Specify VisiumHD platform
resolution = "16um", # REQUIRED for VisiumHD
qc_metric = "sum_umi", # or "sum" depending on your colData
samples = "sample_id",
buffer_width_um = 100, # VisiumHD-specific parameter
mad_threshold = 2.5,
edge_threshold = 0.75,
name = "edge_artifact"
)
# Step 3: Classification Phase - CRITICAL: Scale min_spots for VisiumHD resolution
# For 16µm bins, use ~6-10× the Standard Visium threshold
min_spots_16um <- 30 * (55 / 16)^2 # ≈ 354 bins
spe_hd16_classified <- classifyEdgeArtifacts(
spe_hd16_detected,
qc_metric = "sum_umi",
min_spots = round(min_spots_16um), # ~350 bins
name = "edge_artifact"
)
# Visualization (same approach as Standard Visium)
table(spe_hd16_classified$edge_artifact_classification)
# This is a pseudo-example demonstrating VisiumHD 8µm workflow
# Assumes you have loaded a VisiumHD 8µm SpatialExperiment object as 'spe_hd8'
# Step 1: QC metrics
spe_hd8 <- addPerCellQCMetrics(spe_hd8)
# Step 2: Detection Phase
spe_hd8_detected <- detectEdgeArtifacts(
spe_hd8,
platform = "visiumhd", # Specify VisiumHD platform
resolution = "8um", # REQUIRED: Specify 8µm resolution
qc_metric = "sum_umi",
samples = "sample_id",
buffer_width_um = 100, # Buffer zone in micrometers
mad_threshold = 2.5,
edge_threshold = 0.75,
name = "edge_artifact"
)
# Step 3: Classification with 8µm-appropriate threshold
# For 8µm bins, use ~20-40× the Standard Visium threshold
min_spots_8um <- 30 * (55 / 8)^2 # ≈ 1,420 bins
spe_hd8_classified <- classifyEdgeArtifacts(
spe_hd8_detected,
qc_metric = "sum_umi",
min_spots = round(min_spots_8um), # ~1,400 bins
name = "edge_artifact"
)
table(spe_hd8_classified$edge_artifact_classification)Platform-Specific Function Calls:
| Platform | Function Call | Required Parameters |
|---|---|---|
| Standard Visium | detectEdgeArtifacts(..., platform="visium") |
shifted=TRUE |
| VisiumHD 16µm | detectEdgeArtifacts(..., platform="visiumhd", resolution="16um") |
resolution |
| VisiumHD 8µm | detectEdgeArtifacts(..., platform="visiumhd", resolution="8um") |
resolution |
Parameter Recommendations by Platform:
| Parameter | Standard Visium | VisiumHD 16µm | VisiumHD 8µm |
|---|---|---|---|
platform |
"visium" |
"visiumhd" |
"visiumhd" |
resolution |
N/A (not used) | "16um" |
"8um" |
shifted |
TRUE |
N/A (handled internally) | N/A |
buffer_width_um |
N/A |
100 (default) |
100 (default) |
mad_threshold |
1.5-3.0 | 2.0-3.0 | 2.0-3.0 |
min_spots (classify) |
20-40 | 100-200 | 400-800 |
| Grid Type | Hexagonal | Square | Square |
We’ll create a comprehensive visualization showing QC metrics, detection results, and detailed cluster information:
library(SpatialExperiment)
library(patchwork)
plot_data <- as.data.frame(colData(spe_classified))
plot_data <- cbind(plot_data, as.data.frame(spatialCoords(spe_classified)))
plot_data_in_tissue <- plot_data[plot_data$in_tissue, ]
base_theme <- theme_void() +
theme(
plot.title = element_text(size = 12, hjust = 0.5),
legend.position = "right"
)
# Plot 1: UMI Counts
p1 <- ggplot(plot_data_in_tissue, aes(
x = pxl_col_in_fullres, y = pxl_row_in_fullres,
color = log10(sum_umi + 1)
)) +
geom_point(size = 0.5) +
scale_color_viridis_c(name = "log10(UMI)") +
ggtitle("A. UMI Counts") +
base_theme +
coord_fixed()
# Plot 2: Detected Genes
if ("detected" %in% names(plot_data_in_tissue)) {
p2 <- ggplot(plot_data_in_tissue, aes(
x = pxl_col_in_fullres, y = pxl_row_in_fullres,
color = detected
)) +
geom_point(size = 0.5) +
scale_color_viridis_c(name = "Genes", option = "plasma") +
ggtitle("B. Detected Genes") +
base_theme +
coord_fixed()
} else {
p2 <- ggplot() +
theme_void() +
ggtitle("B. Detected Genes (Data N/A)")
}
# Plot 3: Raw Detection (Any Problem Area)
if ("edge_artifact_problem_id" %in% names(plot_data_in_tissue)) {
p3 <- ggplot(plot_data_in_tissue, aes(
x = pxl_col_in_fullres, y = pxl_row_in_fullres,
color = !is.na(edge_artifact_problem_id)
)) +
geom_point(size = 0.5) +
scale_color_manual(
values = c("FALSE" = "lightgray", "TRUE" = "red"),
name = "Problem?"
) +
ggtitle("C. Raw Detection (All Problem Areas)") +
base_theme +
coord_fixed()
} else {
p3 <- ggplot() +
theme_void() +
ggtitle("C. Raw Detection (Data N/A)")
}
# Plot 4: Cluster IDs
if ("edge_artifact_problem_id" %in% names(plot_data_in_tissue)) {
plot_data_in_tissue$cluster_display <- NA
has_cluster <- !is.na(plot_data_in_tissue$edge_artifact_problem_id)
plot_data_in_tissue$cluster_display[has_cluster] <-
plot_data_in_tissue$edge_artifact_problem_id[has_cluster]
n_clusters <- length(unique(plot_data_in_tissue$cluster_display[has_cluster]))
p4 <- ggplot(plot_data_in_tissue, aes(
x = pxl_col_in_fullres, y = pxl_row_in_fullres,
color = cluster_display
)) +
geom_point(size = 0.5) +
scale_color_discrete(name = "Cluster ID", na.value = "lightgray") +
ggtitle(paste0("D. Problem Area Clusters (n=", n_clusters, ")")) +
base_theme +
coord_fixed() +
theme(
legend.key.size = unit(0.3, "cm"),
legend.text = element_text(size = 8)
)
} else {
p4 <- ggplot() +
theme_void() +
ggtitle("D. Cluster IDs (Data N/A)")
}
# Plot 5: Edge vs Interior Separation
if ("edge_artifact_edge" %in% names(plot_data_in_tissue)) {
plot_data_in_tissue$artifact_type <- "Normal"
# Mark edge artifacts
plot_data_in_tissue$artifact_type[plot_data_in_tissue$edge_artifact_edge] <- "Edge Artifact"
# Mark interior problem areas (not edges)
interior_mask <- !is.na(plot_data_in_tissue$edge_artifact_problem_id) &
!plot_data_in_tissue$edge_artifact_edge
plot_data_in_tissue$artifact_type[interior_mask] <- "Interior Problem"
p5 <- ggplot(plot_data_in_tissue, aes(
x = pxl_col_in_fullres, y = pxl_row_in_fullres,
color = artifact_type
)) +
geom_point(size = 0.5) +
scale_color_manual(
values = c(
"Normal" = "lightgray",
"Edge Artifact" = "red",
"Interior Problem" = "blue"
),
name = "Artifact Type"
) +
ggtitle("E. Edge vs Interior Distinction") +
base_theme +
coord_fixed()
} else {
p5 <- ggplot() +
theme_void() +
ggtitle("E. Edge vs Interior (Data N/A)")
}
# Plot 6: Final Hierarchical Classification
if ("edge_artifact_classification" %in% names(plot_data_in_tissue)) {
p6 <- ggplot(plot_data_in_tissue, aes(
x = pxl_col_in_fullres, y = pxl_row_in_fullres,
color = edge_artifact_classification
)) +
geom_point(size = 0.5) +
scale_color_manual(
values = c(
"not_artifact" = "lightgray",
"large_edge_artifact" = "red",
"small_edge_artifact" = "orange",
"large_interior_artifact" = "blue",
"small_interior_artifact" = "cyan"
),
name = "Final Class",
na.value = "grey50"
) +
ggtitle("F. Final Hierarchical Classification") +
base_theme +
coord_fixed()
} else {
p6 <- ggplot() +
theme_void() +
ggtitle("F. Final Classification (Data N/A)")
}
# Combine all plots in a 3x2 layout
(p1 | p2) / (p3 | p4) / (p5 | p6)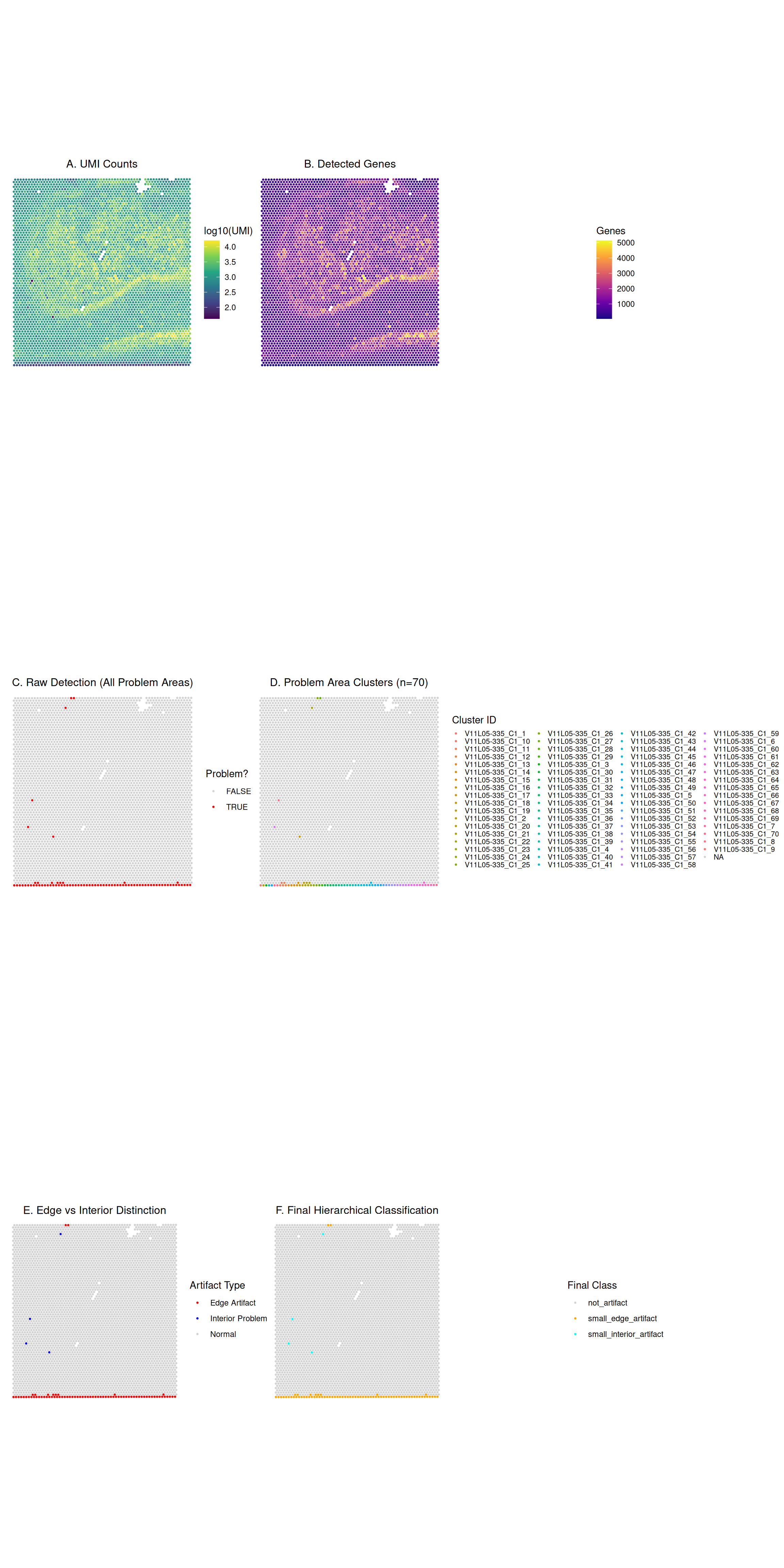
Let’s examine the enhanced classification system:
cat("--- Final Classification Summary (`edge_artifact_classification`) ---\n")
#> --- Final Classification Summary (`edge_artifact_classification`) ---
# Use the 'spe_classified' object we created in the step above
final_summary <- table(spe_classified$edge_artifact_classification)
print(final_summary)
#>
#> not_artifact small_edge_artifact small_interior_artifact
#> 4887 74 4
final_pct <- round(100 * final_summary / sum(final_summary), 2)
final_df <- data.frame(
Classification = names(final_summary),
Count = as.numeric(final_summary),
Percentage = as.numeric(final_pct)
)
print(final_df)
#> Classification Count Percentage
#> 1 not_artifact 4887 98.43
#> 2 small_edge_artifact 74 1.49
#> 3 small_interior_artifact 4 0.08
cat("\n\n")
cat("--- Raw Edge Detection Summary (`edge_artifact_edge`) ---\n")
#> --- Raw Edge Detection Summary (`edge_artifact_edge`) ---
# Use the 'spe_classified' object and the new column name
edge_summary <- table(spe_classified$edge_artifact_edge)
print(edge_summary)
#>
#> FALSE TRUE
#> 4891 74
edge_pct <- round(100 * edge_summary / sum(edge_summary), 2)
edge_df <- data.frame(
Flagged_As_Edge = names(edge_summary),
Count = as.numeric(edge_summary),
Percentage = as.numeric(edge_pct)
)
print(edge_df)
#> Flagged_As_Edge Count Percentage
#> 1 FALSE 4891 98.51
#> 2 TRUE 74 1.49Finally, let’s validate that flagged spots have lower quality metrics:
in_tissue_data <- spe_classified[, spe_classified$in_tissue]
qc_data <- data.frame(
sum_umi = in_tissue_data$sum_umi,
detected_genes = in_tissue_data$detected,
flagged = in_tissue_data$edge_artifact_edge
)
# --- Calculate Median Differences ---
flagged_umi <- median(qc_data$sum_umi[qc_data$flagged], na.rm = TRUE)
nonflagged_umi <- median(qc_data$sum_umi[!qc_data$flagged], na.rm = TRUE)
flagged_gene <- median(qc_data$detected_genes[qc_data$flagged], na.rm = TRUE)
nonflagged_gene <- median(qc_data$detected_genes[!qc_data$flagged], na.rm = TRUE)
cat("QC Validation Results (comparing raw edge detection flag):\n")
#> QC Validation Results (comparing raw edge detection flag):
cat("Flagged spots - Median UMI:", round(flagged_umi), "\n")
#> Flagged spots - Median UMI: 120
cat("Non-flagged spots - Median UMI:", round(nonflagged_umi), "\n")
#> Non-flagged spots - Median UMI: 1784
cat("UMI difference:", round(nonflagged_umi - flagged_umi), "\n\n")
#> UMI difference: 1664
cat("Flagged spots - Median Detected Genes:", round(flagged_gene), "\n")
#> Flagged spots - Median Detected Genes: 106
cat("Non-flagged spots - Median Detected Genes:", round(nonflagged_gene), "\n")
#> Non-flagged spots - Median Detected Genes: 1019
cat("Gene difference:", round(nonflagged_gene - flagged_gene), "\n")
#> Gene difference: 912
qc_data$flag_status <- ifelse(qc_data$flagged, "Flagged (Raw Edge)", "Non-Flagged")
validation_plot <- ggplot(qc_data, aes(x = flag_status, y = log10(sum_umi + 1), fill = flag_status)) +
geom_boxplot() +
scale_fill_manual(values = c("Flagged (Raw Edge)" = "lightcoral", "Non-Flagged" = "lightblue")) +
labs(
title = "QC Validation: UMI Counts in Raw Edge vs Non-Edge Spots",
x = "Raw Edge Detection Status", y = "log10(UMI Count + 1)"
) +
theme_minimal()
print(validation_plot)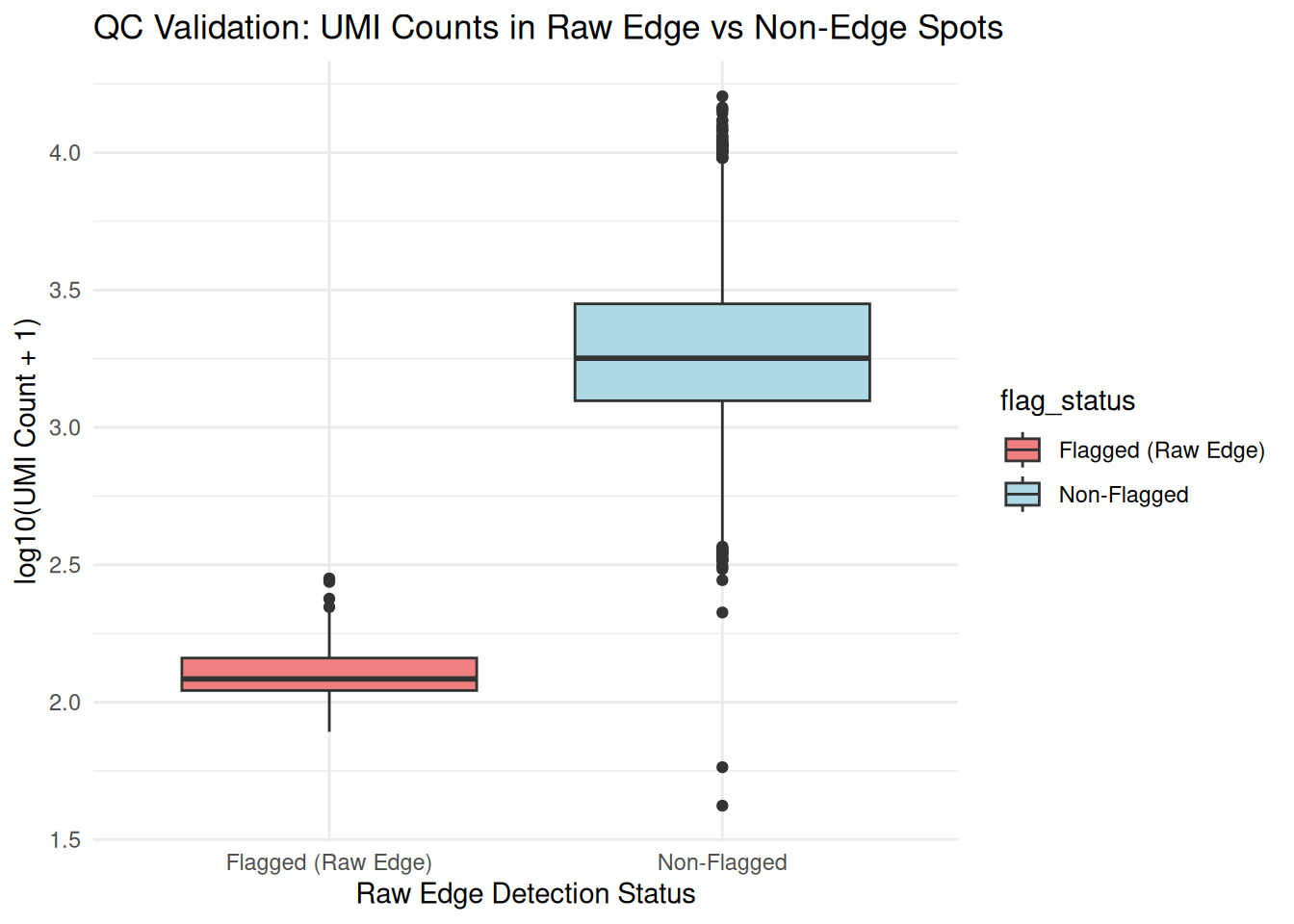
Based on the new classifications, users can make informed decisions about filtering. For example, you might decide to remove all edge artifacts (both large and small) while keeping interior artifacts for further review.
Here’s how you can filter the SpatialExperiment object to remove all spots classified as both "large_edge_artifact" or "small_edge_artifact":
if ("edge_artifact_classification" %in% names(colData(spe_classified))) {
spots_to_keep <- !spe_classified$edge_artifact_classification %in%
c("large_edge_artifact", "small_edge_artifact")
spe_filtered <- spe_classified[, spots_to_keep]
cat("Original number of spots:", ncol(spe_classified), "\n")
cat("Number of spots after filtering:", ncol(spe_filtered), "\n")
} else {
cat("Classification column not found. Filtering step skipped.\n")
}
#> Original number of spots: 4965
#> Number of spots after filtering: 4891
plot_data_before <- as.data.frame(colData(spe_classified))
coords_mat <- spatialCoords(spe_classified)
plot_data_before <- cbind(plot_data_before, as.data.frame(coords_mat))
plot_data_before_in_tissue <- plot_data_before[plot_data_before$in_tissue, ]
plot_data_after <- as.data.frame(colData(spe_filtered))
if (ncol(spe_filtered) > 0) {
coords_mat_after <- spatialCoords(spe_filtered)
plot_data_after <- cbind(plot_data_after, as.data.frame(coords_mat_after))
}
base_theme <- theme_void() +
theme(
plot.title = element_text(size = 12, hjust = 0.5),
legend.position = "right"
)
p1_umi_before <- ggplot(plot_data_before_in_tissue, aes(x = pxl_col_in_fullres, y = pxl_row_in_fullres, color = log10(sum_umi + 1))) +
geom_point(size = 0.5) +
scale_color_viridis_c(name = "log10(UMI)") +
ggtitle("UMI Counts (Before Filtering)") +
base_theme +
coord_fixed()
if (ncol(spe_filtered) > 0) {
p2_umi_after <- ggplot(plot_data_after, aes(x = pxl_col_in_fullres, y = pxl_row_in_fullres, color = log10(sum_umi + 1))) +
geom_point(size = 0.5) +
scale_color_viridis_c(name = "log10(UMI)") +
ggtitle("UMI Counts (After Filtering)") +
base_theme +
coord_fixed()
} else {
p2_umi_after <- ggplot() +
theme_void() +
ggtitle("UMI Counts (After Filtering - No Spots)")
}
p3_class_before <- ggplot(plot_data_before_in_tissue, aes(x = pxl_col_in_fullres, y = pxl_row_in_fullres, color = edge_artifact_classification)) +
geom_point(size = 0.5) +
scale_color_manual(
values = c(
"not_artifact" = "lightgray",
"large_edge_artifact" = "red",
"small_edge_artifact" = "orange",
"large_interior_artifact" = "blue",
"small_interior_artifact" = "cyan"
),
name = "Final Class",
na.value = "grey50",
drop = FALSE
) +
ggtitle("Classification (Before Filtering)") +
base_theme +
coord_fixed()
if (ncol(spe_filtered) > 0) {
p4_class_after <- ggplot(plot_data_after, aes(x = pxl_col_in_fullres, y = pxl_row_in_fullres, color = edge_artifact_classification)) +
geom_point(size = 0.5) +
scale_color_manual(
values = c(
"not_artifact" = "lightgray",
"large_edge_artifact" = "red",
"small_edge_artifact" = "orange",
"large_interior_artifact" = "blue",
"small_interior_artifact" = "cyan"
),
name = "Final Class",
na.value = "grey50",
drop = FALSE
) +
ggtitle("Classification (After Filtering)") +
base_theme +
coord_fixed()
} else {
p4_class_after <- ggplot() +
theme_void() +
ggtitle("Classification (After Filtering - No Spots)")
}
combined_filtering_plot_2x2 <- (p1_umi_before | p2_umi_after) / (p3_class_before | p4_class_after)
print(combined_filtering_plot_2x2)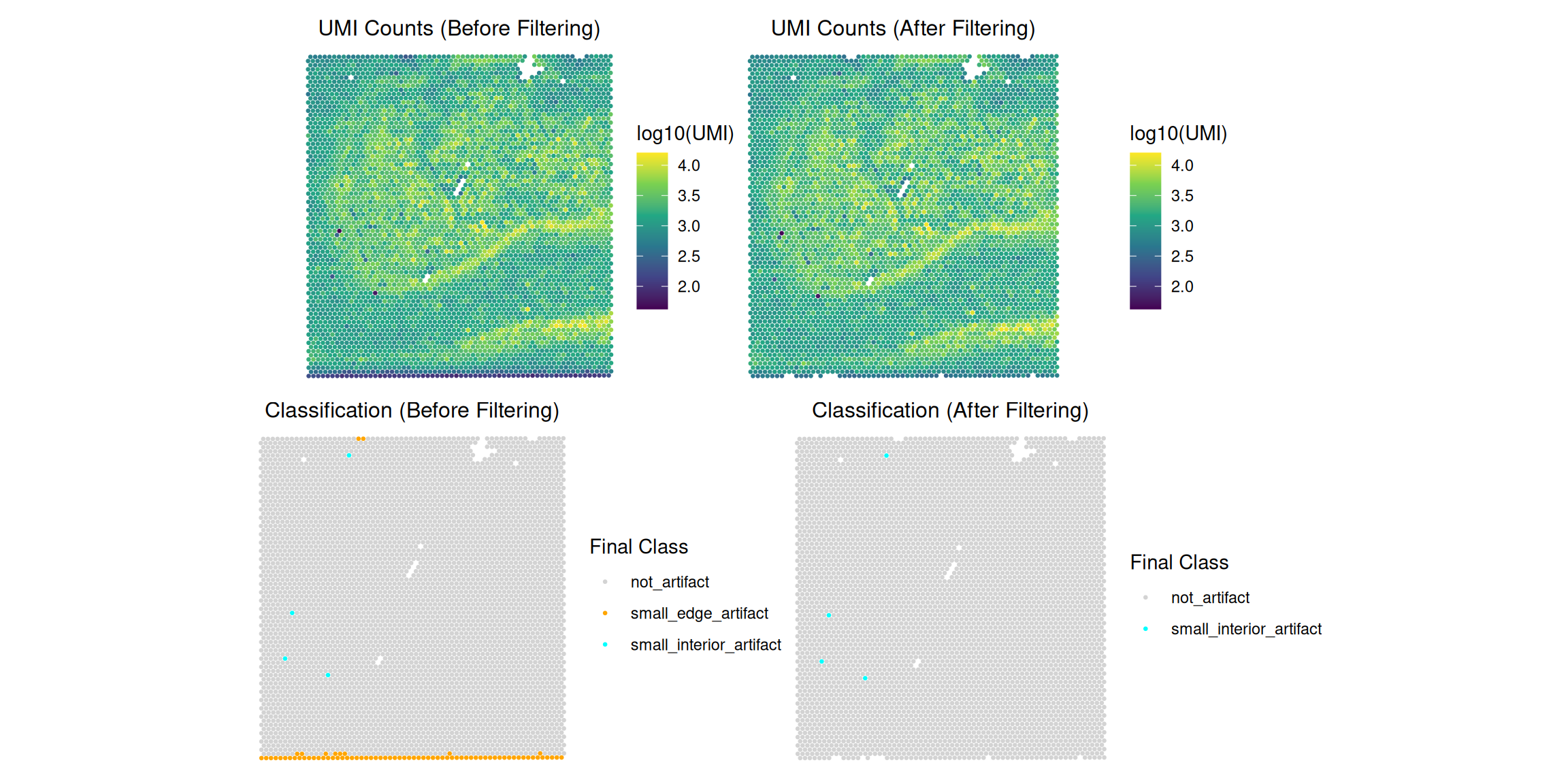
This vignette demonstrated the SpatialArtifacts workflow across multiple spatial transcriptomics platforms. Specifically, it showed:
Key Takeaways for Multi-Platform Usage:
Grid Structure: Standard Visium uses hexagonal grids (shifted = TRUE), while VisiumHD uses square grids (shifted = FALSE)
Critical Parameter Scaling: The min_spots threshold in classifyEdgeArtifacts() must scale with spatial resolution:
min_spots_HD = min_spots_visium × (55 / bin_size)²
Morphological Framework: The same detection logic works across platforms, automatically adapting to different grid arrangements
Physical Consistency: Scaled parameters ensure that “large” and “small” artifact classifications represent equivalent physical sizes regardless of platform
Overall, SpatialArtifacts provides a unified, platform-agnostic framework for detecting and classifying spatial artifacts, enabling consistent quality control across the evolving spatial transcriptomics technology landscape.
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats4 stats graphics grDevices utils datasets methods
#> [8] base
#>
#> other attached packages:
#> [1] dplyr_1.2.0 patchwork_1.3.2
#> [3] ggplot2_4.0.2 SpatialArtifacts_0.99.0
#> [5] SpatialExperiment_1.20.0 SingleCellExperiment_1.32.0
#> [7] SummarizedExperiment_1.40.0 Biobase_2.70.0
#> [9] GenomicRanges_1.62.1 Seqinfo_1.0.0
#> [11] IRanges_2.44.0 S4Vectors_0.48.0
#> [13] BiocGenerics_0.56.0 generics_0.1.4
#> [15] MatrixGenerics_1.22.0 matrixStats_1.5.0
#>
#> loaded via a namespace (and not attached):
#> [1] SparseArray_1.10.8 scuttle_1.20.0 lattice_0.22-7
#> [4] digest_0.6.39 magrittr_2.0.4 evaluate_1.0.5
#> [7] grid_4.5.2 RColorBrewer_1.1-3 fastmap_1.2.0
#> [10] jsonlite_2.0.0 Matrix_1.7-4 viridisLite_0.4.3
#> [13] scales_1.4.0 codetools_0.2-20 abind_1.4-8
#> [16] cli_3.6.5 rlang_1.1.7 XVector_0.50.0
#> [19] withr_3.0.2 DelayedArray_0.36.0 yaml_2.3.12
#> [22] S4Arrays_1.10.1 tools_4.5.2 beachmat_2.26.0
#> [25] parallel_4.5.2 BiocParallel_1.44.0 vctrs_0.7.1
#> [28] R6_2.6.1 lifecycle_1.0.5 magick_2.9.1
#> [31] pkgconfig_2.0.3 terra_1.8-93 pillar_1.11.1
#> [34] gtable_0.3.6 glue_1.8.0 Rcpp_1.1.1
#> [37] xfun_0.56 tibble_3.3.1 tidyselect_1.2.1
#> [40] knitr_1.51 farver_2.1.2 rjson_0.2.23
#> [43] htmltools_0.5.9 labeling_0.4.3 rmarkdown_2.30
#> [46] compiler_4.5.2 S7_0.2.1